
SPECTROstar Nano
Absorbance plate reader with cuvette port
Cell-based assays and cell culture in general are often considered a separate discipline in life sciences. However, in reality there is hardly a discipline in which cell-based assays have not already found their use and the applications continue to grow. Find out more about the different types of cell-based assays available and why cell-based assays are relevant to such a wide range of disciplines.
 Dr Ann-Cathrin Volz
Dr Ann-Cathrin Volz
In contrast to a biochemical assay, in which recombinant proteins or cell lysates are analysed, a cell-based assay measures different parameters in living or fixed cells, like apoptosis or functional gene expression (figure 1). The use of cells results in a great benefit: cells constitute a comprehensive unit, which enables functional analysis and represents the behaviour and reaction of whole living organisms much better than biochemical assays1.
Cell-based assays are used to understand gene, protein or whole cellular functions and the regulatory mechanism controlling these, to screen for potential inhibitors or inducers of biological processes in drug development or to localise an effect or event within the cell.
During the last decades, cell-based assays have evolved from simple 2D layers of adherent monocultures to multicellular assays in a 3D setup². While some approaches require the use of tags to enable the readout with absorbance, fluorescence or luminescence measurements, some allow label-free evaluation.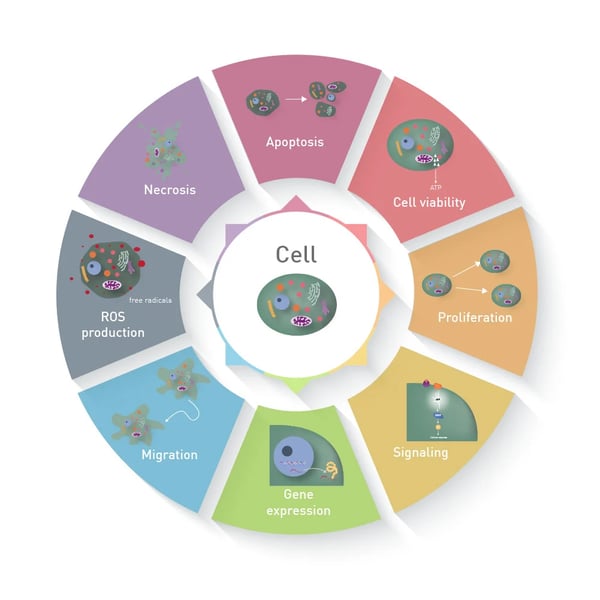 What are cell-based assays used for?
What are cell-based assays used for?
A cell-based assay can be used in basic life science research as well as in drug discovery for screening purposes. In the biomedical field, the use is typically limited to a low to medium sample throughput, whereas in drug screening, cell-based assays are often used in high-throughput procedures.
Life science research
In basic life science research, cell-based assays aim to unravel all mechanisms on which biochemical processes are based – in both the physiological and the pathological state. Furthermore, cells are used as building blocks in biomanufacturing of sophisticated cell-based constructs, mimicking the in vivo situation as closely as possible. These can then in turn be used for test purposes.
Drug discovery
Cell-based assays are used in various stages of drug development. It is a prerequisite to test a drug candidate in defined cell-based in vitro experiments before moving on to live animals or later stage clinical trials. The central question in drug discovery naturally revolves around the biological activity of the test substances. This application note highlights the use of a Resazurin cell viability assay to screen snake venom libraries for a potential apoptotic activity against cancer cell lines. This ‘potency’ of drug products can also be tested with a cell-based assay during drug development. Cell-based assays can be used for the initial ‘hit finding’ to identify a drug with confirmed activity against a biological target. This step of drug discovery depends on knowledge of complex biology, comprehensive screening libraries and innovative drug screening platforms.
Furthermore, an appropriately adapted cell-based assay can of course also be used for lead optimisation during drug development. In drug development, it is important to exclude general adverse reactions from the screened drugs such as a cytotoxic effect. Drug developers use cell-based assays mainly to quantify cytotoxicity and cell death caused by tested compounds. Screened drugs are cycled through a series of complementary cell-based assays in drug discovery campaigns to maximise potency and selectivity while minimising toxicity.
In addition to the evaluation of the biological activity of a drug, cell-based assays can also be used for more detailed investigations during the drug development, allowing the mechanisms of action (MOA) to be unravelled more precisely.
Finally, it is not sufficient to proof the biological activity of a drug. It must also be ensured that there are no off-target interactions. Many drugs fail in the late stage of clinical trials during drug development due to unanticipated side effects in more complex systems. A cell-based assay can’t prevent this completely, but they can help to identify problematic drugs early in drug development campaigns. Therefore, drug targets are validated in a second round of disease-relevant cell-based assays during drug discovery.
The greatest benefit of using cell-based assays is obvious: In contrast to simple biochemical assays, they better reflect the complexity of biological systems, as they do not only contain an isolated receptor or enzyme but all organelles of a cell, organised in a functional cellular unit. This complexity allows to accurately model cellular behaviour in real-life and thereby brings more physiological relevance to in vitro assays and higher reliability e.g., to the identification of potentially effective drugs. Furthermore, cell-based assays provide the opportunity to evaluate different properties of substances at the same time.
A cell-based assay can be used to measure the cellular behaviour over time (up to months) and to understand the influence of environmental factors. Advanced systems for analysing living cells can provide necessary and unique insights into dynamic life processes.
Since cell-based assays can also be miniaturized and evaluated using multiplexing, they are also increasingly used in high-throughput screening (HTS). Many compounds can be tested in parallel under different conditions, in small test volumes, thereby circumventing multi-step screening methods³.
The bottom line is, cell-based assays save a lot of resources, time and money not only in drug development. That is why their use is so popular in various fields.
A cell-based assay can either be based on isolated primary cells or performed with immortalised cell lines. Cell lines do not only exhibit lower demands on cell culture conditions, but also provide more robust results due to smaller heterogeneity in the population. However, the risk that the obtained results may differ from the in vivo model is much lower when using primary cells. These are, however, not easy to get and handle4. The most common cell-based assays can be divided into the following main categories:
Cell viability assays
Cell viability describes the ability of a cell population to live and is regularly expressed as percentage. Essential cellular functions are assessed to estimate the viability of a population. Cell viability assays can be based either on metabolic activity, the cell’s reduction potential or cellular ATP production. The cell’s reduction potential is regularly measured via resazurin or tetrazolium-based assays like MTT or WST. ATP production is most frequently measured with kits using the very ATP as fuel to produce a luminescent signal (figure 2). This AppNote comprises a comparison of different viability assays in terms of lower limit of detection and user-friendliness. Using sophisticated solutions like BMG LABTECH’s Atmospheric Control Unit (ACU), long term culture and accompanying evaluation of cellular reaction in real-time are made possible as shown in this AppNote, studying cell viability and cytotoxicity over 72h.
Proliferation assays
Cell proliferation and cell viability are often used synonymously. However, viable and metabolically active cells do not necessarily have to show cell proliferation. Assays which solely report on cell proliferation are e.g. based on DNA replication and use labelled nucleosides like BrdU (5-Bromo-2’-Deoxyuridine), which is incorporated into DNA instead of thymidine. Other cell proliferation assays are based on the constitutive expression of a fluorescent protein. Such reporter genes, can for example also be used to study transfection efficiency, e.g. using the gene sequence for Green Fluorescent Protein (GFP) introduced into the cell.
Cytotoxicity assays
When cell viability decreases, markers for cytotoxicity or cell death increase. Cytotoxicity describes the ability of a compound or organism to harm cells, ultimately leading to cell death. Cytotoxicity assays make use of events that happen during cell death such as loss of membrane integrity, activation of cell death-inducing enzymes called caspases or phenotypic changes on the cell surface. Some even allow to differentiate between apoptosis and necrosis. The loss of cell membrane activity can for instance be detected with the RealTime-GloTM Annexin V assay employing a fluorescent DNA detection reagent, which is only able to reach the DNA when cell membrane is compromised during necrosis. At the same time the assay detects the translocation of phosphatidylserine from the inner to the outer membrane during apoptosis with a luminescence-based readout. This additionally allows to differentiate between apoptosis and necrosis.
ROS assays
Reactive oxygen species are unstable molecules, which can damage DNA, RNA and proteins, and even cause cell death but at the same time play an important role in regulating cellular signalling and gene expression. Detecting ROS levels can be difficult as they are often present at low concentrations and have very short half-lives. 2',7'-dichlorodihydrofluorescein diacetate (H2DCFDA) is the most common reagent used for measuring ROS. MitoSOX™ Red selectively detects superoxide generated in mitochondria. The fluorescent probe roGFP (redox sensitive green fluorescent protein) has different excitation wavelengths dependent on its redox state and is genetically encoded in the cells genome for ROS measurement. Chemiluminescent probes like L-012 emit light when transitioning to their oxidised state.
Signalling/cytokine responses
Several methods exist for monitoring receptor-mediated activation of cellular signalling, including those that directly measure secondary messengers such as cAMP or intracellular calcium or those which use response elements upstream of a primary reporter to measure transcriptional activation. In this AppNote, CRISPR/Cas9 was used to introduce the gene sequence of a protein, functioning as biosensor reporting on β-arrestin signalling in living cells to analyse G-Protein Coupled Receptor activation (figure 3).
Migration assays
Cellular migration is a key procedure in many processes including tissue formation, immune defence but also cancer progression5. Cell-based assays intending to measure migration include scratch- or wound healing assays. Here, a confluent monolayer is scratched and migration of cells into the scratched area is monitored. Furthermore, transwell assays monitor the migration of cells from one compartment to the other. The movement of cells, is most easily monitored with cells expressing fluorescent proteins or luciferases or by the use of cell dyes as demonstrated in this OrisTM Cell Migration assay.
Protein degrader assays
Cell-based assays can be used in microplates to study all the individual steps involved in targeted protein degradation. Researchers can choose from different options for cell-based protein degrader assays . For PROTACs and molecular glues , techniques like AlphaScreen® , Time-Resolved Fluorescence Resonance Energy Transfer (TR-FRET) or luminescence methods like NanoBRET™ can be performed in microplates to measure the interaction between target and ligase. These techniques can also be used to study targeted protein degradation that arises from interactions between known degrons and ligases.
BMG LABTECH microplate readers offer various dedicated solutions for the application of cell-based assays:
The Atmospheric Control Unit (ACU) enables you to fully and independently regulate both O2 and CO2 gas levels within the microplate reader according to the requirements of your cell-based assay. From standard cell culture conditions to hypoxic assays, the ACU is the ideal solution for live cell-based assays.
Reading directly from the bottom of the well comes with benefits, especially for cell-based assays with fluorescence and in rare assays also luminescence readout, since both the distance between the cell layer and the detector and the effect of the cell culture medium are minimised.
The signal intensity of cell-based samples is often hard to predict – especially for kinetic assays in which the maximum signal builds up over a lapse of time. This makes detection of all samples with the same gain problematic. The Enhanced Dynamic Range (EDR) function provides the ability to measure bright and dim signals without the need for gain adjustment and without manual intervention.
Software-controlled reagent injectors enable reagent delivery in your cell-based assay at desired injection timing, injection speed and delivery volume to any plate format from 6-384 wells. This enables for instance the performance of a calcium assay, requiring fast kinetics.
BMG LABTECH’s various shaking options are not only needed to keep suspension cells in motion, but for instance also to increase oxygen uptake and maintain aerobic conditions in a cell culture. The microplate reader’s incubator additionally allows to control air temperature up to 45° or 60/65°C, with extended incubation. Thereby the optimal temperature conditions can be established in cell-based assays – individually for each cell type.
Even when using cell lines, one cell is rarely the same as another. BMG LABTECH’s well scanning options allow to compensate for cellular heterogeneity within cell-based assays by averaging single measurements at different positions in the well. Using the matrix scan options even provides you with a local resolution of the signal throughout the well. See how the different scanning options can influence the data variability in our HowTo Note: How to reduce data variability in heterogenous cell samples.
Cellular autofluorescence can significantly affect the measurement results or even make them completely unusable. Since this autofluorescence in cell based assays is mainly associated to the green/yellow wavelength range (figure 4), the usage of red-shifted dyes is a promising tool to reduce adverse effects from the cells or their culture medium in cell-based assays. BMG LABTECH offers dedicated red-shifted PMTs which offer higher sensitivity in the red wavelength range up to 900nm.
The CRISPR/Cas9 technology has revolutionised the study of genes and their functions as it allows very accurate and easy genome editing. CRISPR greatly simplifies the development of more complex disease-relevant cell-based assays and supports researchers in focussing their attention on more complex questions and heterogenous diseases with more complex and biologically relevant assays.
Not only the complexity of the individual cell used but also of the complete culture system is increasing. Next to the translation to physiological 3D environments and the inclusion of multiple cell types to create co-cultures and study mutual interdependencies, cell-based assays are also introduced into dynamic systems in bioreactors and miniaturised in so-called microfluidic chips.
In his testimonial, Building a bridge between organoid research and drug discovery, Elad Katz from Navigate Precision Biology discusses the establishment of an organoid platform in 384-well format and how the PHERAstar FSX has been instrumental to developing accurate tests.

Absorbance plate reader with cuvette port
Powerful and most sensitive HTS plate reader
Most flexible Plate Reader for Assay Development
Upgradeable single and multi-mode microplate reader series
Flexible microplate reader with simplified workflows
Learn about applications for bacterial metabolism on a microplate reader.
Life in the depths of the ocean operates under extreme conditions. Find out how proteins from deep-sea luminescent organisms are useful for measurements on microplate readers.
Second messengers play a pivotal role in signal transduction events in cells. But how do you measure these small, transiently lived molecules and how can microplate readers help?
NanoBRET is used to analyse binding events, signaling pathways and receptor trafficking in live cells and has significantly expanded the range and applications of BRET assays.
Read here about the various threats associated with mycoplasma contamination and find out about methods to prevent, detect, and eliminate mycoplasma in your cell culture.