
PHERAstar FSX
Powerful and most sensitive HTS plate reader
NanoBRETTM (Nanoluciferase Bioluminescence Resonance Energy Transfer) is a detection technique used to analyse binding events, signaling pathways and receptor trafficking in live cells. Through the adoption of the Nanoluciferase, it has significantly expanded the range and applications of BRET assays.
 Dr Tobias Pusterla
Dr Tobias Pusterla
Tobias Pusterla’s scientific background spans veterinary biotechnology, cancer cell biology, and the molecular mechanisms underlying inflammation‑driven tumorigenesis. After graduating in Veterinary Biotechnology at the University of Milan, Italy, he worked in mouse mutagenesis before completing a Ph.D. in Cellular and Molecular Biology through a joint program between the Open University of London, UK and the San Raffaele Scientific Institute, Milan, Italy. He later conducted postdoctoral research at the German Cancer Research Center (DKFZ) in Heidelberg, Germany, focusing on tumor biology, the tumor microenvironment, and the role of chronic inflammation in cancer development. His scientific work has contributed to understanding how damage‑associated molecular signals drive immune activation, cell migration, inflammation, and tumorigenesis, helping to clarify fundamental pathways linking cellular stress responses to physiological and pathological outcomes. After more than 13 years of research experience, he joined BMG LABTECH in 2013. Here, he oversees global marketing activities, including the creation of scientific content and the coordination of application support.
Luminescence is one of the most common techniques used in the life sciences to analyse and quantify biological and biochemical processes and interactions. It mainly relies on the two enzymes Firefly (Fluc) and Renilla luciferases (Rluc) to biochemically generate light in the presence of their substrate and does not require an excitation light.
Fluc and Rluc have also been used for Bioluminescence Resonance Energy Transfer (BRET) experiments. Similarly to FRET and TR-FRET, BRET relies on resonance energy transfer between two chromophores. However, in BRET the energy transfer takes place between a luciferase donor and a fluorophore acceptor.
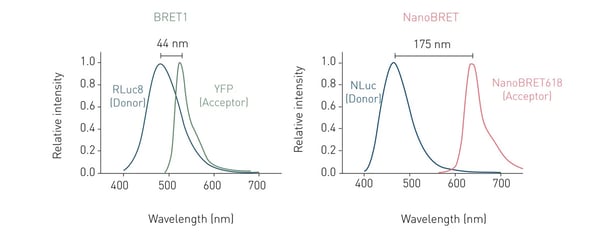
For BRET to take place, the emission spectrum of the donor luciferase needs to overlap with the excitation spectrum of the fluorophore acceptor. Additionally, donor and acceptor molecules need to be in close proximity (typically 1-10 nm), as the resonance energy transfer decreases with distance (fig. 1).
Typically, donor and acceptor are tagged to proteins of interest to monitor protein-protein proximity and interaction in live cells. The output, known as BRET ratio, is the intensity of the acceptor divided by the intensity of the donor. 1
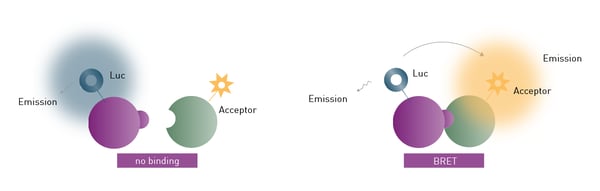
There are two main requirements for BRET assays. First, the donor and acceptor tags must not interfere with the function, configuration or cellular localisation of the target proteins. Second, a microplate reader with multi-chromatic luminescence detection and the capability to measure live cells in real-time is required for its quantification. 2
The most commonly used BRET setup (also known as BRET1), utilises Rluc as energy donor and Yellow Fluorescent Protein (YFP) as acceptor. This pairing achieves an efficient spectral overlap and hence enables resonance energy transfer. However, the spectral proximity between Rluc and YFP generates high background noise, negatively affecting sensitivity and dynamic range (fig. 2).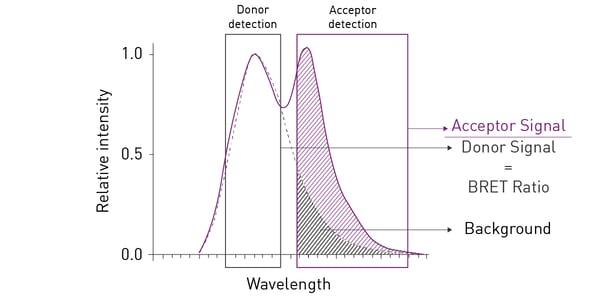 Consequently, BRET2 was developed to increase spectral separation between donor and acceptor and decrease background noise. BRET2 uses a substrate analogue, DeepBlueCTM, which shifts the Rluc emission peak to shorter wavelengths from about 475 nm with coelenterazine as substrate, to around 395 nm. As donor, GFP2 or GFP10 (Green Fluorescent Protein analogues) are used as they have excitation and emission peaks at 395 and 510 nm, respectively. An example of a BRET2 assay can be found in the application note Sequential or simultaneous emission detection for BRET assays.
Consequently, BRET2 was developed to increase spectral separation between donor and acceptor and decrease background noise. BRET2 uses a substrate analogue, DeepBlueCTM, which shifts the Rluc emission peak to shorter wavelengths from about 475 nm with coelenterazine as substrate, to around 395 nm. As donor, GFP2 or GFP10 (Green Fluorescent Protein analogues) are used as they have excitation and emission peaks at 395 and 510 nm, respectively. An example of a BRET2 assay can be found in the application note Sequential or simultaneous emission detection for BRET assays.
Although the large Stokes shift in BRET2 improves spectral resolution and reduces background noise, the emission intensities are much lower than with BRET1, mainly because of a poor quantum yield and luminescence stability. 1
NanoBRET is the latest addition and was developed to overcome the previous weaknesses.
Table 1: Comparison of different BRET set-ups
| Name | Donor luciferase | Substrate | Donor emission peak | Acceptor fluorophore | Acceptor emission |
| BRET 1 | Renilla (36kDa) | Coelenterazine | 475 nm |
YFP GFP (Venus) |
515-560 nm 510-540 nm |
| BRET 2 | Renilla (36kDa) | DeepBlueC | 395 nm |
GFP2, GFP10 NanoBRET 618 TMR |
500-540 nm 550-675 nm 550-600 nm |
| NanoBRET | NanoLuc | Furimazine | 460 nm |
AlexaFluor 633 Venus Bodipy (BY630) |
618 550-675 nm 515-575 nm 600-700 nm |
NanoBRET is based on the Nanoluciferase (NanoLuc®; Nluc) with furimazine as substrate and was developed by Promega. Nanoluciferase is the result of the optimisation of a multi-component enzyme present in Oplophorus gracilirostris, a deep-sea shrimp (fig. 3).
 The result of Promega´s optimisation is a small (19 kDa) enzyme with greatly enhanced luminescence (around 150 times the intensity of Fluc or Rluc; fig. 4) and stability (> 2 half-life). Nluc was initially used in gene expression studies as a reporter gene. 3
The result of Promega´s optimisation is a small (19 kDa) enzyme with greatly enhanced luminescence (around 150 times the intensity of Fluc or Rluc; fig. 4) and stability (> 2 half-life). Nluc was initially used in gene expression studies as a reporter gene. 3
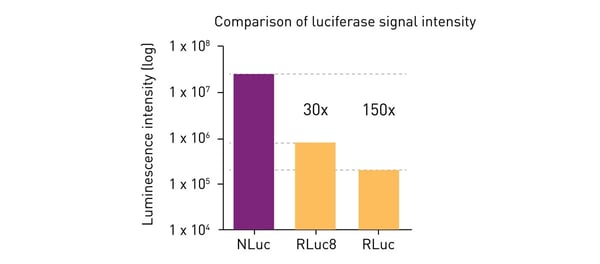
The small size and stronger emission of Nluc led to its rapid uptake in life science and pharmaceutical research, gradually replacing the previously used luciferases, especially where greater sensitivity was required.
The adoption of Nanoluciferase in BRET assays resulted in the creation of a new methodology called NanoBRET. 4 Here, Nluc's stronger luminescence and smaller size offer several advantages over BRET1 and BRET2.
Compared to Rluc (used for BRET), Nluc has a slightly blue-shifted signal with an emission peak at about 460 nm and a narrower spectral range. In NanoBRET, Promega combined the more effective Nluc donor to a new acceptor, HaloTag®.
HaloTag (HT) is a two-component tag system made of a small protein (33 kDa) that can be genetically fused to a protein of interest and a chloroalkane ligand that can be covalently bound to a fluorophore or other molecules. The best acceptor performance was achieved by matching HT with a red-shifted fluorophore with emission peak at about 620 nm, called NanoBRET 618. This provides a spectral separation from the Nluc of around 170 nm, leading to decreased background noise and improved sensitivity (fig. 5). 4
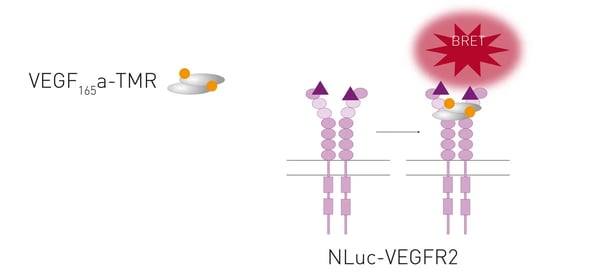
TAMRA is a commonly used fluorophore with an excitation peak of around 556 nm and an emission peak at about 579 nm. NanoBRET was used to monitor binding of the vascular endothelial growth factor (VEGF) tagged with TAMRA to the receptor VEGFR2. 5 This approach is further discussed in our application note NanoBRET assay quantitatively evaluates VEGF binding to the VEGFR2 in real-time in living cells (fig. 6).
Another versatile option is the use of BODIPY fluorophores. BODIPY fluorophores are hydrophobic and have hence good solubility in several solvents, but not in aqueous solutions. In addition, they are relatively unaffected by polarity or pH changes and multiple BODIPY colour variations are commercially available. In our application note NanoBRET assay for monitoring of ligand binding to GPCRs in live cells, GPCR ligand binding was monitored by using BY630 as acceptor. Similarly to BODIPY, Alexa FluorTM dyes are available in different colours and have been frequently used in FRET and TR-FRET assays.
Other fluorophores available as NanoBRET acceptors in the green range are 4-nitro-7-aminobenzofurazan (NBD) and Venus. The latter is a GFP variant, whereas NBD is smaller than TAMRA and BODIPY molecules and exhibits increased fluorescence in hydrophobic environments such as the plasma membrane.
The advantages of NanoBRET and the use of Nluc over BRET1 and BRET2 methods include:
NanoBRET assays are typically run in microplates. Besides microscopic applications, the major part of NanoBRET assays are run in microplates and detected with microplate readers. For this, a luminescence microplate reader with multi-chromatic luminescence detection is required. Moreover, since NanoBRET is mainly used in live cell-based assays for kinetic detection, temperature and atmospheric control should be strongly considered.
NanoBRET assays require wavelength selection as two signals (the donor and the acceptor) are emitted by one sample. Accordingly, plate readers need to be able to select and separately quantify the light emitted by Nluc and the light coming from the acceptor fluorophore to calculate the NanoBRET ratio (fig. 8).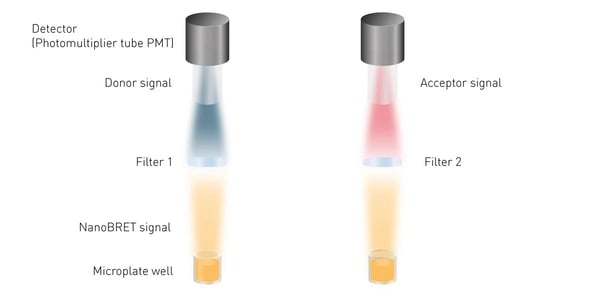
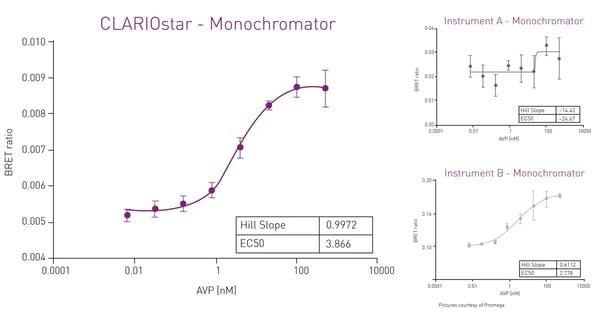
Filter-based plate readers are typically recommended for wavelength selection in NanoBRET assays. Conventional grating-based monochromators play a subordinate role because of their limited sensitivity in the acceptor channel that is mainly due to scattering effects and narrow bandwidths. However, a monochromator technology using Linear Variable Filters (LVF) as available in the CLARIOstar Plus reader provides the sensitivity needed for luminescence detection. An LVF-based monochromator exhibits filter-like light transmission and provides bandwidths up to 100 nm. This assures best performance also in NanoBRET assays (fig. 9).
 The detection of the two emission wavelengths can be performed sequentially or simultaneously, depending on the plate reader set-up. Typically, readers measure the two signals one after the other. The PHERAstar FSX however can detect the two signals simultaneously, as it is equipped with Simultaneous Dual Emission and a pair of assay-matched detectors (PMTs). This feature halves detection time and reduces data variability.
The detection of the two emission wavelengths can be performed sequentially or simultaneously, depending on the plate reader set-up. Typically, readers measure the two signals one after the other. The PHERAstar FSX however can detect the two signals simultaneously, as it is equipped with Simultaneous Dual Emission and a pair of assay-matched detectors (PMTs). This feature halves detection time and reduces data variability.
BMG LABTECH plate readers for NanoBRET detection include the Omega series, VANTAstar, CLARIOstar Plus and PHERAstar FSX.
Since its introduction, NanoBRET has been used to analyse molecular processes with a multitude of applications mainly focussed on protein interaction studies and real-time monitoring of binding events in living cells, as typically cells do not have to be lysed.
Ligand binding studies have been very hard to carry out with BRET1 and BRET2 set-ups mainly because of the large size of the Rluc and Fluc luciferases. In contrast, thanks to the small size of the Nluc, NanoBRET was successfully used in ligand binding and has numerous advantages over traditional radioligand binding assays. For instance, GPCRs were N′-terminally tagged with Nluc as shown in the application note NanoBRET assay for monitoring of ligand binding to GPCRs in live cells (fig. 10). 2
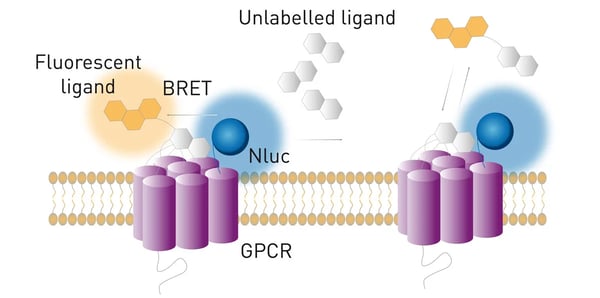 Moreover, the strong intensity of Nluc enables the characterisation of low affinity ligands. Importantly, NanoBRET enables the investigation of ligand binding kinetics in real time in live cells, expanding the possibilities of ligand binding studies.
Moreover, the strong intensity of Nluc enables the characterisation of low affinity ligands. Importantly, NanoBRET enables the investigation of ligand binding kinetics in real time in live cells, expanding the possibilities of ligand binding studies.
In this scientific talk, Thomas Machleidt from the Promega corporation discusses the advantages of NanoBRET over conventional BRET and the advantages of the CLARIOstar plate reader. Moreover, he shows a few case studies of protein-protein interaction assays in live cells.

NanoBRET can also be used in pharmacology for instance for real-time receptor profiling in live cells. This includes ligand binding, secondary messenger coupling, recruitment of scaffolding proteins and internalisation and trafficking. This is addressed in the scientific talk “Real-time profiling of receptor pharmacology”.

In this interview Professor Kevin Pfleger further discusses his work on BRET-based ligand binding and how he uses the PHERAstar and CLARIOstar plate readers for this purpose.

The CRISPR/Cas9 system has revolutionised genome engineering. The strong signal of Nluc in combination with the precise editing capabilities of CRISPR/Cas9 enable more physiologically relevant studies in cells. This combination was firstly reported in 2017 and is further discussed in our application note CRISPR/Cas9 genome-edited cells express nanoBRET-donor that monitors protein interaction and trafficking. 7
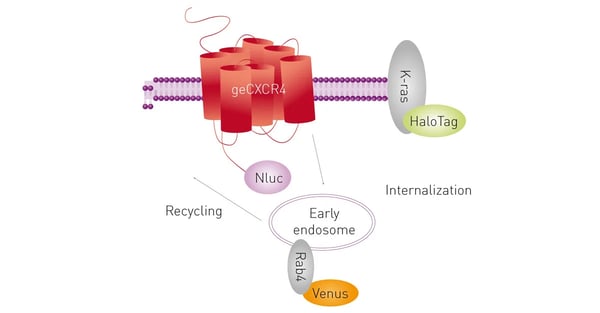
Here, NanoBRET was used to analyse the recruitment of genome-edited β-arrestin2 and compared to the recruitment of exogenously-expressed (transfected) β-arrestin2 to multiple GPCRs. Moreover, a NanoBRET multiplex assay with two acceptors was used to study CXCL12-induced receptor internalisation and trafficking (fig.11).
Interestingly, a NanoBRET signal could also be achieved using endogenous expression of Nluc-tagged proteins through CRISPR/Cas9-mediated homology-directed repair.
Another example of NanoBRET-based ligand-receptor interaction studies is further discussed in our application note NanoBRET assay quantitatively evaluates VEGF binding to the VEGFR2 in real-time in living cells, where binding of the vascular endothelial growth factor (VEGF) to the receptor VEGFR2 was monitored.5 Additonal material on NanoBRET ligand-receptor assays can be found in the application note: “Novel nanoBRET-Based Biosensor Enables Comprehensive Screening of Estrogenic Endocrine-Disrupting Chemicals (eEDCs) in Live Cells”.
In the following interview Laura Kilpatrick, Chloe Peach and Mark White, authors of the mentioned scientific papers and the application notes discuss their work and the advantages of the BMG LABTECH microplate readers.

For a general overview on interaction assays in drug discovery, in this scientific talk our application specialists Anne and Martin highlight how advanced detection technologies such as NanoBRET and TR-FRET can be used in high-throughput drug screening.

Powerful and most sensitive HTS plate reader
Most flexible Plate Reader for Assay Development
Upgradeable single and multi-mode microplate reader series
Flexible microplate reader with simplified workflows
Learn about applications for bacterial metabolism on a microplate reader.
Life in the depths of the ocean operates under extreme conditions. Find out how proteins from deep-sea luminescent organisms are useful for measurements on microplate readers.
Second messengers play a pivotal role in signal transduction events in cells. But how do you measure these small, transiently lived molecules and how can microplate readers help?
Find out about the different types of cell-based assays and why they have become an indispensable tool in a such a broad variety of disciplines in this blog article.
Read here about the various threats associated with mycoplasma contamination and find out about methods to prevent, detect, and eliminate mycoplasma in your cell culture.