
PHERAstar FSX
Powerful and most sensitive HTS plate reader
Measurements of binding kinetics are crucial for drug research. Find out how binding kinetics work and which approaches can be used to study them on microplate readers.
Molecular interactions lie at the heart of biology. In binding kinetic experiments, the interaction of two or more biomolecules with each other is studied over time and is the foundation of pharmacological theory. But what is the motivation behind these experiments?
 Dr Martin Mangold
Dr Martin Mangold
Dr. Martin Mangold works as an Applications Specialist at BMG LABTECH headquarters in Ortenberg, Germany. He studied biology with a focus on biochemistry and cell biology at the University of Bonn before specializing in pharmaceutical sciences and drug interactions in his doctoral studies. During his time in the pharmaceutical department, Dr. Mangold gained expertise in protein sciences, binding and interaction studies, and enzyme kinetics as part of an interdisciplinary team of chemists, pharmacists and biologists. Since 2021, Dr. Mangold has been part of the BMG LABTECH team where he authors application notes, performs training courses and supports scientific customers.
Binding interactions are crucial events that rely on the proximity of two biomolecules to each other as well as their affinity. Without binding interactions there would be no enzymatic activity, no receptor signalling, no cellular transport and no DNA transcription and translation. In short, life would not be possible.
Misregulated binding events can cause various diseases. This loss of function can express itself in various ways including for example the accumulation of protein aggregates (found in Alzheimer´s and prion disease) or in unwanted enzymatic activity that can lead to irregular degradation processes (found for example in autoimmune diseases). Here, studying binding kinetics is a viable tool for better understanding the underlying mechanisms of a disease and identifying suitable targets for treatment.
Accordingly, binding kinetics (specifically receptor-ligand kinetics) are an important aspect of drug research. Drugs are often specifically designed to target and eventually inhibit specific binding interactions. Binding kinetics are for example used in drug discovery studies for the screening of large libraries of small molecules for potential therapeutics. These drugs must be as potent and specific as possible to prevent undesirable side effects. Kinetic parameters that can be determined in binding kinetics, such as binding affinity, are crucial tools for this purpose and provide information about different aspects of the interaction.
Proteins are widespread in biological systems and are often involved in the formation and breakdown of molecular complexes. They are essential for many molecular events including enzyme-catalyzed reactions, signal transduction and the transport and storage of substances. The kinetics of protein binding, which include the association and dissociation rates, are therefore crucial for understanding the efficiency and specificity of these processes. Understanding the kinetics of protein binding helps in drug discovery and development to target specific protein interactions.
Common targets for drugs comprise kinases and GPCRs which are the focus of many studies 1, 2, but also other protein-protein interactions regarding the proteasome or other proteases. Find out more about the importance of binding kinetics in the pharmaceutical industry in this scientific talk by Prof. Steven Charlton: The kinetics of drug binding - Why it is important?

Binding interactions are made possible by various forces that allow intermolecular connections such as Van der Waal´s interactions, hydrogen bonding, electrostatic forces and hydrophobic effects. While most interactions are non-covalent and reversible, some binding interactions are based on covalent bonds that are formed between biomolecules. In this scientific talk featuring Martin Redhead, you can learn more about the advantages of kinetic binding methods in contrast to binding affinity analysis at equilibrium.

Independent of the type of binding interaction, it is important to note that binding is not static. The interaction of two biomolecules is comprised of association events according to their association rate and dissociation events dependent on their dissociation rate. Binding changes over time until it reaches equilibrium at which point the concentrations of binding complexes are constant. Binding kinetics can also be used to analyse other factors that can influence the time course of an interaction such as for example temperature, pH, viscosity, or the concentrations of interaction partners.
For binding kinetics, three parameters are particularly important. The association, or on-rate (kon) defines how much of a ligand (L) binds to a biomolecule (B) over a specific time to form the biomolecule ligand complex (BL). The dissociation rate or off-rate (koff) defines the amount of ligand that dissociates from the biomolecule ligand complex over a specific time. Together they form the equilibrium dissociation constant KD of a ligand which is the ratio of dissociation to association events over time. KD indicates the affinity of a ligand to a biomolecule and is an important parameter for binding and drug research: The lower the KD value the higher the affinity of a ligand or drug to its target which means lower ligand concentrations are needed to induce binding. However, since high affinity can be the cause of either a low koff rate or a high kon rate it is important to determine these parameters in binding kinetics to fully understand the binding behaviour of a drug or ligand and choose the right concentration of the drug to administer. For example, drugs with a high kon rate have a faster onset of action, while the effect of drugs with a low koff rates lasts longer since the drug remains bound to its target longer.
Different approaches can be used to study binding kinetics depending on the biomolecules under investigation, desired readout and kinetic parameters.
How are studies of binding kinetics carried out? Below I focus on the four approaches that are used most often to study binding kinetics on microplate readers.
Förster´s Resonance Energy Transfer (FRET) and time-resolved FRET (TR-FRET) are probably the most popular detection modes used for performing binding kinetics. Both are based on energy transfer between two fluorophores. These fluorophores are attached to the two binding partners of a binding interaction. Since they have overlapping emission and excitation spectra, emission of the donor fluorophore leads to the subsequent excitation of the acceptor fluorophore, but only when both fluorophores and binding partners are in close proximity. However, since FRET can be negatively affected by scattered excitation light and autofluorescence in a sample, it can suffer from high background signals. TR-FRET, which uses long-lived lanthanide donor fluorophores, was developed to reduce background signals.
The application note "Binding kinetics: high throughput assay for kinase inhibitors" highlights a microplate-based TR-FRET assay for performing binding kinetics. Here, the assay was used to determine the target binding kinetic rates kon and koff for kinase inhibitors. This was achieved by sequentially performing association and dissociation binding kinetics (Fig. 1) which allowed screening of 270 potential inhibitors against a panel of 40 kinase drug targets in a high-throughput manner. The versatility of TR-FRET assays for performing binding kinetics is highlighted in the application notes analysis of binding kinetics with HTRF or the detection of human tau protein aggregation.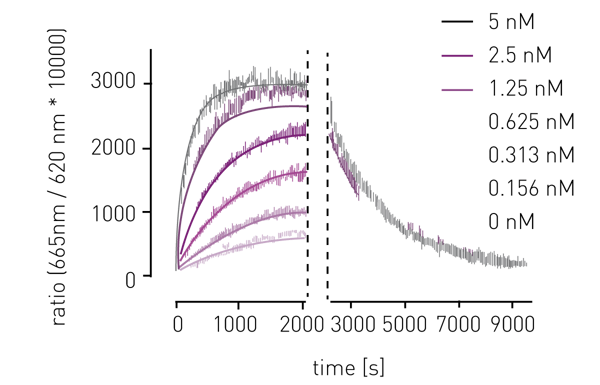
BRET assays are also based on the resonance energy transfer between two interaction partners and can be used to perform binding kinetics. In contrast to FRET, the donor signal in BRET assays is provided by a luciferase which generates emission light by an enzymatic reaction (fig. 2). Accordingly, no additional excitation source must be defined. As with FRET, BRET assays are performed as ratiometric measurements of acceptor and donor signal which allows comparisons to be made between different test runs.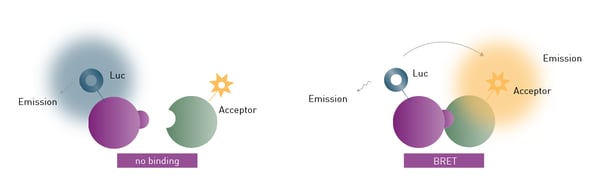 However, the available BRET assay systems BRET1 and BRET2, which differ in substrate and emission characteristics, have some shortcomings regarding sensitivity and background noise. NanoBRET overcomes these limitations since it offers greatly enhanced luminescence signal intensity and stability by using a modified luciferase derived from a deep-sea shrimp species. This luciferase provides an emission signal 100x brighter than conventional firefly or Renilla luciferases.
However, the available BRET assay systems BRET1 and BRET2, which differ in substrate and emission characteristics, have some shortcomings regarding sensitivity and background noise. NanoBRET overcomes these limitations since it offers greatly enhanced luminescence signal intensity and stability by using a modified luciferase derived from a deep-sea shrimp species. This luciferase provides an emission signal 100x brighter than conventional firefly or Renilla luciferases.
In the application note "NanoBRET™ assay quantitatively evaluates VEGF binding to the VEGFR2 in real-time in living cells" , a novel labelling technique was used to study multiple VEGF isoforms and quantitatively evaluate their binding to VEGFR2 in real-time in living cells (Fig.3). In another study, ligand binding to GPCRs was investigated and dissociation constants calculated. Alternatively, BRET assays can also be used to study G-protein dissociation as well as β-arrestin recruitment to GPCRs upon ligand binding. Novel luminescence-based approaches use the NanoBIT® system, which employs a fragmented nanoluciferase attached to two binding partners of interest. Upon binding of the two binding partners, the nanoluciferase fragments form a functional enzyme that can generate a luminescence signal as highlighted in the application note: Identification of androgen-disruptors using a cell-based androgen receptor dimerization assay.
Fluorescence polarization assays are used to measure binding kinetics and are often used to study protein-protein interactions, protein-DNA binding, as well as enzyme activity.
Fluorescence polarization uses small fluorescent ligands which on their own have a rapid tumbling rate. Accordingly, when the polarized excitation light excites the fluorescence ligand, the emission light becomes depolarized. After binding to a larger biomolecule, the tumbling rate of the fluorescent ligand is reduced, and its emission light retains most of its polarization (fig. 4). Binding kinetics are measured by comparing the polarization of unbound fluorescent ligand to bound fluorescent ligand.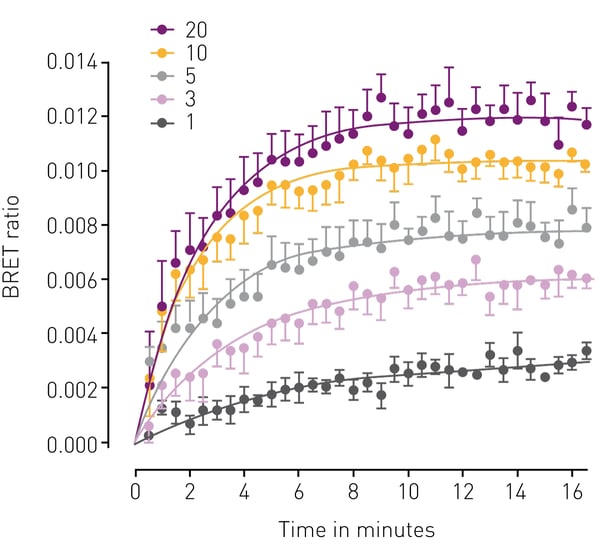 This principle was used in the application note "Protein-ligand binding measurements using fluorescence polarization" to investigate the binding kinetics of human Lys-specific histone demethylase 1 (LSD1) to histone-derived peptides. Here, binding assays for three different peptides were performed, followed by competition experiments with potential competitors. The application note Competition assay using Fluorescence Polarisation to determine the Residence Times for Calcitonin and AMYR agonist, AM833, demonstrates that fluorescence polarization can also be employed to determine association and dissociation rates of ligand receptor interactions. Fluorescence polarization is also suited for high-throughput binding kinetics as highlighted in the application note "Fluorescence polarization based assay suitable for screening H-Prostaglandin D Synthase inhibitors".
This principle was used in the application note "Protein-ligand binding measurements using fluorescence polarization" to investigate the binding kinetics of human Lys-specific histone demethylase 1 (LSD1) to histone-derived peptides. Here, binding assays for three different peptides were performed, followed by competition experiments with potential competitors. The application note Competition assay using Fluorescence Polarisation to determine the Residence Times for Calcitonin and AMYR agonist, AM833, demonstrates that fluorescence polarization can also be employed to determine association and dissociation rates of ligand receptor interactions. Fluorescence polarization is also suited for high-throughput binding kinetics as highlighted in the application note "Fluorescence polarization based assay suitable for screening H-Prostaglandin D Synthase inhibitors".
AlphaScreen® is a bead-based chemistry used to perform binding kinetics. Today AlphaScreen technology includes multiple assay formats, which are mainly used in the life sciences for screening purposes.
AlphaScreen uses specific donor and acceptor beads bound to two interaction partners of interest, respectively. A luminescent light signal is generated when the donor and acceptor beads are in close proximity and energy transfer takes place. The special feature of this method is that the excitation light has a longer wavelength than the measured emission signal. 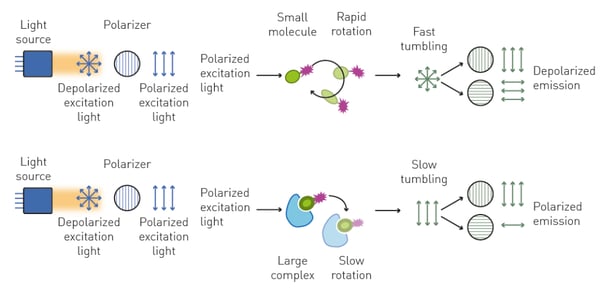 AlphaScreen was employed in the application note "PHERAstar measures AlphaScreen assay to develop selective inhibitors for the human YEATS domains" which highlights an assay that can be used for inhibitor screening. Here, several compound libraries with a total of over forty thousand potential inhibitors were screened against the ENL YEATS domain, yielding a potent and selective small molecule inhibitor.
AlphaScreen was employed in the application note "PHERAstar measures AlphaScreen assay to develop selective inhibitors for the human YEATS domains" which highlights an assay that can be used for inhibitor screening. Here, several compound libraries with a total of over forty thousand potential inhibitors were screened against the ENL YEATS domain, yielding a potent and selective small molecule inhibitor.
Microplate readers are the gold standard for binding kinetics even though methods like surface plasmon resonance or biolayer interferometry have been used in the past. In a microplate format, binding kinetics can be used for high throughput screening of large compound libraries, decreasing the measurement time and need for resources including reagents and personnel.
BMG LABTECH’s multi-mode microplate readers are ideally suited for binding kinetics experiments.
They can be equipped with more than one of the detection modes needed for most binding kinetic assays. The CLARIOstar® Plus or the VANTAstar® offer the ideal solution for flexibility and sensitivity for all of your applications.
However, if you are looking for the most sensitive and fastest detection of binding kinetics possible, PHERAstar® FSX is your go-to reader. The PHERAstar FSX is equipped with special features for this purpose, such as dedicated TRF and Alpha Technology lasers to increase TR-FRET and AlphaScreen assay performance. Furthermore, due to four dedicated photomultiplier tube detectors, the PHERAstar can measure at two distinct emission wavelengths simultaneously. The Simultaneous Dual Emission (SDE) feature reduces measurement time by half and binding kinetic measurements are readily obtained as ratiometric measurements. At the same time, data quality is increased, since the two signals derive from the same excitation event and are less susceptible to other potential influencing variations.
Moreover, the AAS system provides a steady temperature that is unaffected by external environmental changes while the plate reader is operating. This provides improved assay stability for temperature-sensitive reactions such as binding reactions.
In his testimonial, Pioneering Pre-Clinical Drug Discovery with Advanced Assays, Nick Holliday from Excellerate Bioscience and the University of Nottingham discusses the benefits of the PHERAstar FSX for TR-FRET-based kinetic binding analysis.

Powerful and most sensitive HTS plate reader
Most flexible Plate Reader for Assay Development
Flexible microplate reader with simplified workflows
Binding constants quantify the strength of a binding reaction between a biomolecule and its target (ligand). But how do you measure them and what can you do with them?
Receptor-ligand kinetics is the study of the rates at which receptors and ligands interact, bind and dissociate. Learn why these types of measurements are important and how to measure them.
Receptor-ligand interactions are crucial for cell signalling. They are also important for drug discovery. How do microplate readers deliver benefits to both?
Degrons are specific sequences of amino acids or structural motifs within a protein that are important for targeted protein degradation. Find out how microplate readers can advance research into natural and engineered degrons.
Gene reporter assays are sensitive and specific tools to study the regulation of gene expression. Learn about the different options available, their uses, and the benefits of running these types of assays on microplate readers.
Innovation for targeted protein degradation and next-generation degraders is gathering pace. This blog introduces some of the different approaches that act via the lysosome or proteasome.